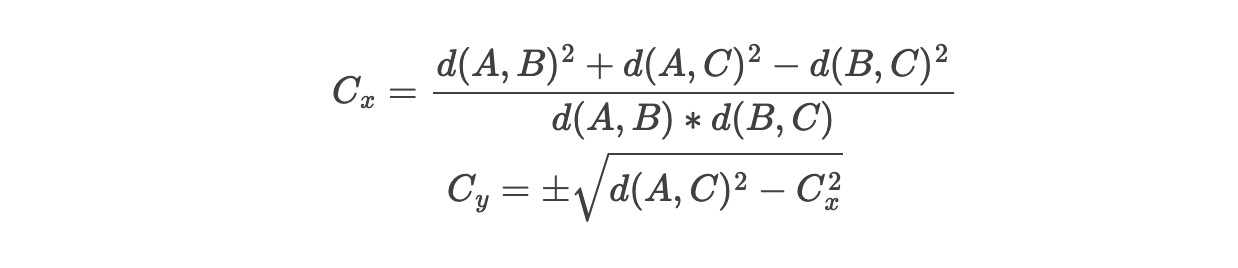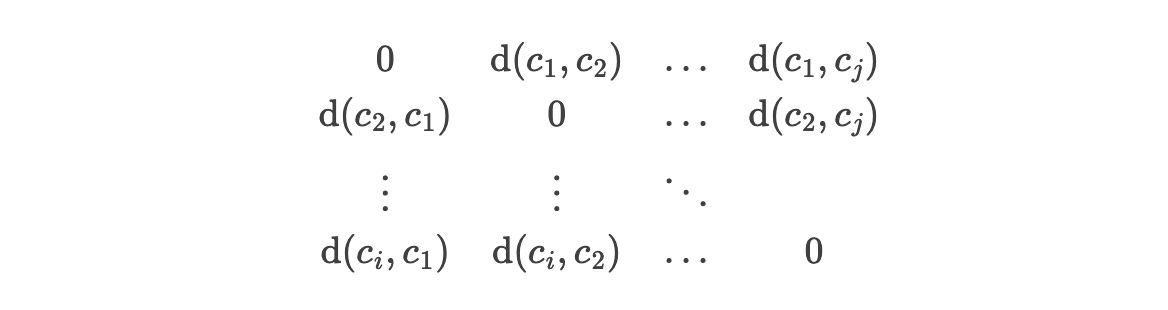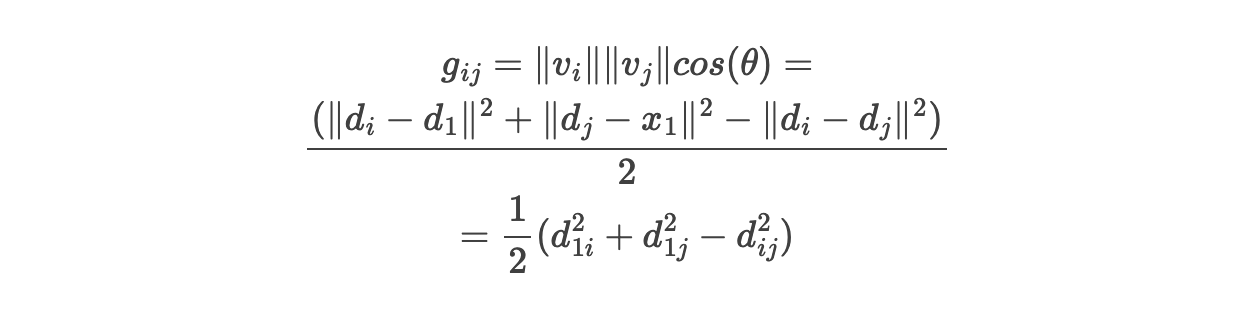If someone gives you a distance matrix, can you generate plausible (x, y) coordinate pairs?
The first thing to realize is that we are dealing with infinite solutions. If we can find a single solution (you can imagine a 2D city map), by repeatedly performing arbitrary rotations / reflections / translations (so-called Euclidean transformations) on the city map, we change the coordinates while preserving the distances of each pair of cities and thus we never run out of novel solutions.
The easy case: three cities
Let’s try generating a solution for three cities. Given cities {A, B, C} and distances d(A, B), d(B, C), d(A,C) it’s easy to notice that a valid solution is the set of vertex coordinates of a triangle with edge lengths corresponding to the distances. You can use a ruler and compass: any line segment A̅B̅ of length d(A, B) generates the first two vertices and the intersection of two arcs centered at A and B with corresponding radiuses d(A, C) and d(B, C) will form the vertex C.
The computational approach for the 3 city problem would work like this: start with A=(0, 0) and B=(0, d(A,B)). The compass intersaction is a system of two Pythagorean theorem equations (origin to C and B to C) and with a few rearrangements we can compute a solution for C (note: flipping the sign on y works too - it’s an example of vertical reflection):
The general case: distance and Gram matrices
What about thousands of cities? We need an N×N matrix where each row is composed of the distances of all pairs of cities, but note that we include the distance from each city to itself too (which is obviously zero):
NxN matrices are called square matrices because count(rows) = count(columns). The zeroes are the distance-to-self entries and they form the diagonal of the matrix. Since we are working with a square matrix of Euclidean distances, we want to check out Euclidean distance matrices. The only nitpick is that Euclidean matrices are supposed to have entries that are distances squared to simplify operations that revolve around the Pythagorean theorem. That’s fine - we can square each entry. Note the symmetry between an entry (i, j) and an entry (j, i): this comes from the fact that the distance from A to B = the distance from B to A.
So now we represent the distances in an Euclidean distance matrix D, but how do we represent the solution — 2D coordinates of each city? We can make an N×2 matrix (call it S) where each row represents the x and y coordinates of each city (i.e. a 2D vector). There isn’t a unique way to map D into S because as we said earlier, there are infinitely many solutions. The dimensions of the input D collapse during the transformation from N×N to N×2 - this is an example of dimensionality reduction.
Relation to Gram matrices & dot products
Now, there are different ways of doing dimensionality reduction, depending on which geometric/statistical property you want to preserve when you reduce the dimensions. The properties mentioned in the Wiki article for Euclidean distance matrices such as symmetry and zero diagonals are useless when it comes to coordinates (our solution matrix isn’t a square matrix at all).
But scroll a bit down, and there’s one striking property, Euclidean distance matrices are related to Gram matrices: square matrices that are describing each pairwise dot product of a set of vectors. A dot product v₁ ⋅ v2 is summing the pairwise products of all their coordinates: v1 ⋅ v2 = v1ₓ × v2ₓ + v1ᵧ × v2ᵧ. We can easily see that a dot product of v to itself v ⋅ v is x² + y². That’s the length of v squared (let’s also use ||v|| to represent the length of the vector, also called magnitude).
The geometric interpretation is that v1 ⋅ v2 = ||v1|| × ||v2|| × cosθ. So the entries entries of a Gram matrix G are of form g(i, j) = ||v_i|| × ||v_j|| × cos(θ), where θ is the angle between v_i and v_j. Thus, by storing pairwise dot products, we indirectly store the length and angle of each vector.
The proven relation between D=d(i, j)² and G=g(i, j) is quite trivial addition and substraction:
Turning a Gram matrix into a set of points
The “Finding a vector realization” section further down is what we need, but it’s a bit convoluted since it also considers complex vectors. In any case, it says several decomposition techniques of G can produce the N×2 final result.
Now, explaining matrix decomposition techniques and performing them by hand will require a lot of practice with matrix multiplication, transposes, eigenvectors, but fortunately we can use any numeric computing language/library to do it for us. I chose Sage (which can execute on Web) and SVD decomposition:
N = 4
# note 1: Sage indices are 0..N-1 instead of 1..N
# so we gotta be careful.
# distance matrix.
# note 2: the given distances are not squared
D = matrix([[ 0, 3, 10, 1 ],
[ 3, 0, 5, 4 ],
[ 10, 5, 0, 11 ],
[ 1, 4, 11, 0 ]
])
# calculate Gram matrix G as follows:
# g_ij = 1/2 * ( (d_0i)^2 + (d_0j)^2 - (d_ij)^2)
G = matrix(N, N, lambda i, j: (D[0, i] ** 2 + D[0, j] ** 2 - D[i, j] ** 2) / 2)
# SVD of a Gram matrix gives us U * S * U^t
# a possible solution is the product U * √S,
# where the x, y coords of each row are the first two columns
U, S, UT = matrix(RDF, G).SVD()
(U * S.apply_map(lambda el: sqrt(el))).matrix_from_columns([0, 1])If you run it on https://sagecell.sagemath.org/ you get a possible solution:
[ 0.0 0.0]
[ -3.92291410062551 2.528288457959308]
[-10.051441221192283 -1.0157568878859065]
[ 0.965029794343825 -0.3021276784799517]You can check whether this is a valid solution by computing the distances. I got +-15% inaccuracies; presumably because of the square roots and SVD approximations.